NASA Early Career Scientist Spotlight featuring Dr. Angel Mojarro
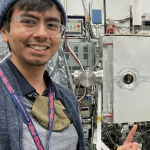

The following spotlight is reposted from: https://science.gsfc.nasa.gov/600/ECSS/Angel-Mojarro.html. Link to Dr. Mojarro’s GSFC biography: https://science.gsfc.nasa.gov/sci/bio/angel.mojarro Early Career Scientist Spotlight Dr. Angel Mojarro (he/him/his) Organic GeochemistAstrochemistry Laboratory (691) What is your research focus? My research is focused on biomarker preservation and the origins of life. In other words, I study how fossils form, how they might persist through […]
QQBlue!

Our wonderful colleagues at Geomark Research shared some Geomark shirts with us after our most recent meetup at the 2022 Gordon Research Conference for Organic Geochemistry. We couldn’t help but take new group photos with them — especially in front of our GC/MS Triple Quad! Thanks, Geomark!
A Semester of BIOmarkers

This year, Summons Lab graduate students Fatima Husain and Angel Mojarro, along with UROP Juliana Drozd, kicked off BIOmarkers, an audio series that archives the oral history of organic geochemistry. Take a listen! Episode 8 – Dr. Katherine Freeman, Part 2 In the eighth episode of BIOmarkers, the audio series that archives the oral history […]
To Mars… and Back!
On November 17th, 2020, Roger Summons and graduate student Fatima Husain were invited to speak during MIT Better World’s Toronto science event. Husain hosted two sessions that discussed the Mars 2020 mission and sample return, while Summons, along with fellow EAPS faculty Tanja Bosak and Benjamin Weiss, shared their expertise as members of the expert […]
Highlighting Women in Geobiology: Dr. Heather Graham

Current Role: I am a research associate with Catholic University of America but I work at NASA’s Goddard Space Flight Center in the Astrobiology Analytical Laboratory. Research Foci: My research focuses on developing tools and techniques for biosignature detection, especially for life that does not share the same biochemistry as life on Earth. What drew […]
Highlighting Women in Geobiology: Dr. Dawn Sumner

Current Role: I am currently a professor at the University of California, Davis, in the Department of Earth and Planetary Sciences. I am also part of the Microbiology Graduate Group at UCD, as well as Chair of the Advisory Board for the Feminist Research Institute at UCD. Research Foci and Initiatives: My main research interests […]
Graduate Student Fatima Husain gives an MIT Chat

This summer, Summons Lab graduate student and Curiosity Correspondent Fatima Husain spoke with students from around the world as part of her MIT Chat. Check out the webinar in which Fatima describes organic geochemistry and biology and her path to it to middle school students.
Highlighting Women in Geobiology at MIT: Dr. Lily Momper

Current Role: I work in the field of environmental consulting for a firm called Exponent. Years in the Summons Lab: I was in the Summons Lab as the W.O. Crosby Postdoctoral Fellow from 2016-2018. Favorite Memory: My favorite MIT-related memories were the Summons lab retreat in 2017 and field work I did in Hamelin Pool, Western Australia (north […]
Highlighting Women in Geobiology at MIT: Dr. Phoebe Cohen

Current Role: I’m an Associate Professor in the Geosciences Department at Williams College, a liberal arts college in the Berkshires. Years at MIT: 2010-2012 Favorite Memory: Wow so many! I got to go to Australia twice while I was at MIT to help create virtual field trips for the NASA Astrobiology grant I was working […]



